Q-FiberMapper with Lesion Load Measures
Q-FiberMapper (QFM) segments major white matter structures in the brain using tractography based on constrained spherical deconvolution (for HARDI) or tensor (for DTI) models.
More information about the Q-FiberMapper tool: Q-FiberMapper Documentation
Then, it uses a lesion mask to compute a set of lesion load measures in each of the segmented fascicles. The lesion load measures are the following:
- Volumetric damage: consists of measuring the relative volume of lesions with respect to total tract volume.
-
Track weighted volumetric damage: In volumetric damage, there is the implicit assumption that all lesion points contribute to bundle damage with equal importance. In this modification, the contribution of the lesion voxels is weighted by the number of streamlines traversing said voxel (taken from the bundle visitation map previously calculated), in order to reflect that a lesion in a point with high density of fibers is probably more disruptive than a lesion at the edge of a bundle where density of fibers might be lessened due to partial volumes.
- Fiber-count damage: When a lesion completely prevents the transmission of signal through the white matter, the fact that a streamline is intersected by a lesion voxel might be more indicative of bundle disruption than a mere percentage of damage volume. To investigate this possibility, a new metric denominated fiber-count damage was considered and computed as the percentage of streamlines that traverse any lesion with respect to the total number of fibers in the considered bundle. A streamline is considered to traverse a lesion when at least 1 of its points lie within a lesion voxel. This value can be modified in the settings of the lesion load measures tool.
-
Length-weighted fiber count damage: In this modification of the previous measure, the computed damage is weighted based on the length of the damaged fibers compared to the sum of the lengths of all the fibers of the fascicle.
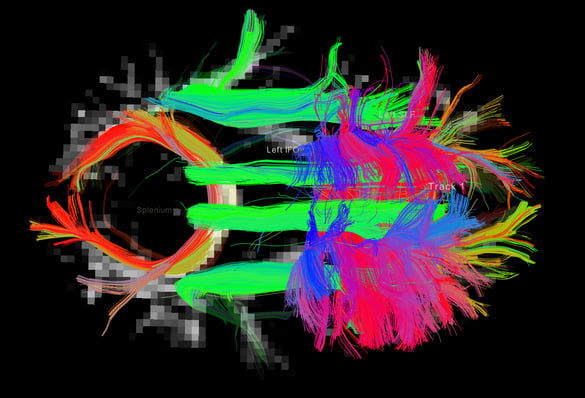
Tract segmentation tool output example:
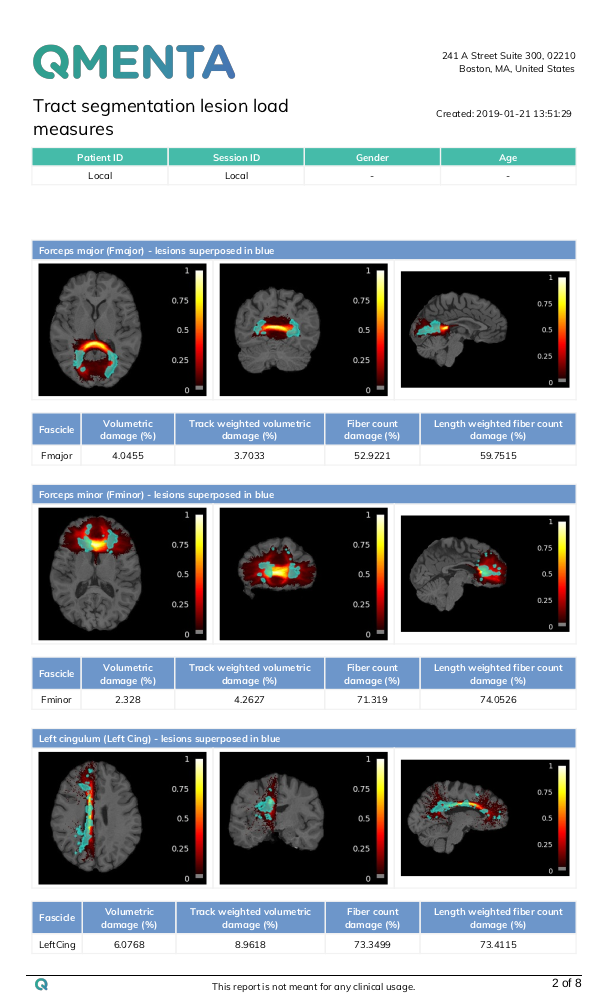 |
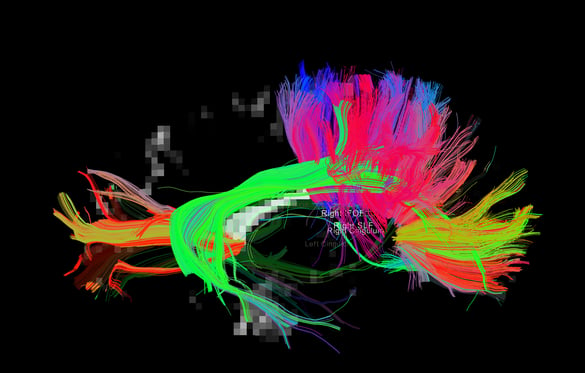 |
 |
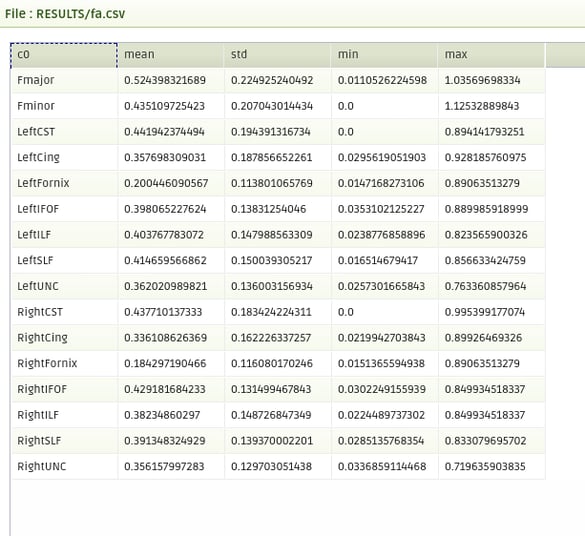 |
The full workflow includes:
-
DICOM files to NIfTI conversion and reorients to standard. The workflow file filter sends the dicom or nifti files required for the workflow to run and the box converts the dicom to niftis, extracts the necessary files (if possible) to do the dmri preprocessing step (eddy) and gradient tables. If nifti files or gradient tables are found in the input, it just does the reorient to standard step and outputs the files.
-
dMRI preprocessing includes denoising, gibbs ringing correction, eddy correction, motion correction, bias field correction, distortion correction (if gradient field maps are available) and ACPC alignment.
-
Gets the brain mask and computes the tensor from the diffusion image, plus derives different scalar measures: FA, AD, MD, RD, Cl, Cs, Cp.
-
Runs ants cortical volumetry script over T1 image to extract the tissue segmentation file.
-
Applies an atlas to the segmented mask of the cortex (gray matter + subcortical gray matter + brainstem + cerebellum)
-
Align the diffusion image to the T1
-
Estimate the diffusion model. Does nothing if the data is DTI. Runs constrained spherical deconvolution model.
-
Perform probabilistic or deterministic tractography over the diffusion model.
-
Extracts fascicles from tractography database, creates probabilistic maps of fascicle membership
-
Outputs per fascicle mean diffusion metric fractional anisotropy (FA), mean/radial/axial diffusivity (MD, RD, AD), coefficient of linearity/planarity/sphericity (Cl, Cp, Cs)
-
Joins the results in a pdf report with orthographic-projections of the brain with the fascicles in overlay.
-
Computes per fascicle a set of lesion load measures.
-
Creates a report with the lesion load measures and orthographic-projections of the brain with the fascicles in overlay.
-
Combines de results of the two report tools.
Required inputs:
-
T1: anatomical 3D image
Isotropic resolution recommended
Must be labeled as 'T1' modality -
dMRI: diffusion 4D image (DTI or HARDI).
Isotropic resolution recommended.
Must be labeled as 'DTI' or 'HARDI' modality. -
Gradients: gradient table (optional if dMRI is DICOM).
MRtrix format supported: file must have '.b' extension and be labeled as 'gradient_table'.
FSL format supported: files must have '.bvec' and '.bval' extensions and be labeled 'bvec' and 'bval'. -
Lesion mask: binary mask containing the lesions.
Must be in the same space as the T1 image.
Must be tagged as 'mask'.
Minimum input requirements:
- Image must contain full brain DWI dataset.
- For optimal result reliability, an isotropic resolution is highly recommended
- T1 recommended resolution: 1 mm isotropic.
- T1 minimum reliable resolution: 2mm isotropic.
- dMRI recommended resolution: 2 mm isotropic.
- dMRI minimum reliable resolution: 3 mm isotropic.
- T1 recommended resolution: 1 mm isotropic.
- If DTI image must have at least 7 volumes (1 b0 and 6 diffusion volumes).
- If HARDI image must have at least 21 volumes (1 b0 and 20 diffusion volumes).
- For optimal preprocessing reliability, 1 b0 image is recommended every 10 gradient volumes.
Optional inputs:
- GFMs: Gradient field-maps, recommended (magnitude image and phase image).
Same resolution and FOV as dMRI image recommended.
Files must be labeled 'gfm_magnitude' and 'gfm_phase' respectively.
Files must additionally be labeled 'dti' or 'hardi' depending on dMRI modality label.
Settings
DCM2NII:
- Preferred DICOM to NIfTI conversion tool (drop-down selection):
The selected tool will be tried first to convert DICOM to NIfTI. If the conversion fails, the other options will be tried sequentially until a successful conversion.- DCM2niix (Default)
- Mrtrix
- DCM2nii
- diffunpack
- MRIConvert
ANTS:
- Compute thickness map (checkbox)
Perform additional cortical thickness on T1 (default unchecked)
PREPROCESSING:
- Do denoising (checkbox)
(default checked) - Do Gibb ringing correction (checkbox)
(default checked) - Do Eddy (checkbox)
(default checked) - Do Fugue (checkbox)
(default checked) - Use manual diffusion acquisition parameters (checkbox)
If checked uses the parameters input manually in the following settings (Default unchecked). - Echo Spacing (s) (float)
The time between echoes in an echo planar image (EPI) (Default 0.0003ms) - Echo Factor (integer)
Also called Echo Train Length (ETL), the number of echoes acquired in a single repetition time (TR) (default 100) - Acceleration Factor (integer)
The number of EPI lines acquired in parallel (Default 1) - Echo Difference (s) (float)
(only needed if GFMs are provided), the difference in Echo Time (ET) between GFM acquisitions. (default 0.00246 for SIEMENS acquisitions) - AC-PC alignment (checkbox)
(default checked) - Resample dMRI image (checkbox)
(default checked) - dMRI isotropic resampling resolution (mm) (float)
Resamples the dMRI image to the indicated isotropic resolution in mm. (Default 2 mm)
MASK_ALIGNMENT:
- Compute non-linear registration EPI to T1 (FA input required) (checkbox)
(default checked)
ATLAS:
- Brain atlas for parcellation (drop-down selection)
The selected tool will be tried first to convert DICOM to NIfTI. If the conversion fails, the other options will be tried sequentially until a successful conversion.- DKT40 (Mindboggle-101) (default)
- AAL
MODEL:
- Tracking algorithm type (drop-down selection)
- Constrained Spherical Deconvolution (CSD) (default)
- Tensor
- Multi-tissue constrained spherical deconvolution
TRACTOGRAPHY:
- Max. spherical harmonic order (drop-down menu):
- 4
- 6 (default)
- 8
- 10
- Tracking algorithm type
- DET deterministic
- PROB probabilistic
- Number of streamlines (integer)
Number of streamlines to generate (default 1M) - Step size (mm) (0 for adaptative default) (float)
(default 0) - Use Anatomical constrained tractography (checkbox)
Only works when Morphology results for ACT are passed as input (default checked) -
Stopping angle (º) [0 for adaptive default] (integer)
(default 0) - Min. streamline length (mm) [0 for adaptive default] (integer)
(default 0) - Max. streamline length (mm) [0 for adaptive default] (integer)
(default 0) - Min. FA/FOD cutoff (float)
(default 0)
LESION LOAD MEASURES:
-
Threshold for volumetric damage computation (decimal)
- default: 0.05
- recommended under 0.1
- default: 0.05
- Minimum # of points in a streamline that must fall within a lesion voxel to be considered damaged (integer)
- default: 1
Output container files
We explain the output files for each box. The modalities are shown in colored boxes and the tags are shown with outlined boxes. Each element represents a relevant file from the container, more might be present but they are not as important.
DCM2NII
- T1 : T1 NIfTI file.
- HARDI / DTI : dMRI NIfTI file.
- MASK : Lesion mask NIfTI file.
- HARDI / DTI:
- HARDI / DTI ACQ_PARAMS: acquisition parameter file for eddy preprocessing automatically extracted from dicom file.
- HARDI / DTI BVAL: BVAL file FSL format.
- HARDI / DTI BVEC: BVEC file in FSL format.
- HARDI / DTI GRADIENT_TABLE: gradient table in mrtrix format.
- HARDI / DTI INDEX: index file for eddy preprocessing automatically extracted from dicom file.
ANTS
- T1 HEAD Full T1 head image.
- T1 BRAIN Skull stripped T1.
- T1 MASK Binary skull stripped brain mask .
- TISSUE_SEGMENTATION 6-tissue segmentation image (1: CSF, 2: gray matter, 3: white matter, 4: deep gray matter, 5: brainstem, 6: cerebellum).
- CSV Volumetry information for ICV and each segmented tissue.
PREPROCESSING
- HARDI / DTI : dMRI NIfTI file processed with the steps selected in the settings.
- GRADIENT_TABLE: gradient table in mrtrix format.
- AVERAGE_B0S: Average b0s from the processed dMRI image.
- REPORT: PDF report of the box.
MEASURES
- TENSOR: Diffusion tensor data.
- AVERAGE_B0S: Average b0s from the processed dMRI image.
- MASK: dMRI binary brain mask.
FA
- SCALAR AD: AD scalar file.
- SCALAR CL: Cl scalar file.
- SCALAR CP: Cp scalar file.
- SCALAR CS: Cs scalar file.
- SCALAR FA: FA scalar file.
- SCALAR MD: MD scalar file.
- SCALAR RD: RD scalar file.
- REPORT : PDF report of the box.
ATLAS
- ATLAS_INFO : Text file with atlas information.
- LABELS : Labeled cortical brain.
- REPORT : PDF report of the box.
- Vectorized data folder
- VECTOR : Volumetry information in a vector format.
MASK_ALIGNMENT
- HARDI / DTI : dMRI NIfTI file aligned to the anatomical image.
- SCALAR AD: AD scalar file aligned to the anatomical image.
- SCALAR CL: Cl scalar file aligned to the anatomical image.
- SCALAR CP: Cp scalar file aligned to the anatomical image.
- SCALAR CS: Cs scalar file aligned to the anatomical image.
- SCALAR FA: FA scalar file aligned to the anatomical image.
- SCALAR MD: MD scalar file aligned to the anatomical image.
- SCALAR RD: RD scalar file aligned to the anatomical image.
- ACT : ACT volume (5tt format) for connectome.
- AVERAGE_B0S: Average b0s from the processed dMRI image aligned to the anatomical image.
- GRADIENT_TABLE: gradient table in mrtrix format.
- LABELS : Labeled cortical brain for connectome.
- MASK: dMRI binary brain mask aligned to the anatomical image.
- TISSUE_SEGMENTATION 6-tissue segmentation image for connectome.
MODEL
- CSD / DTI / In the multi-tissue case we have:
- MSMT WM
- MSMT GM
- MSMT CSF
- RESPONSE
TRACTOGRAPHY
- TRACKS : Full tractography.
- TRACKSDOWNSAMPLED : Tractography for visualization.
TRACT SEGMENTATION TOOL
- BUNDLES : Segmented probability bundles.
- TRACKS: Segmented fascicles as trk files.
TST REPORT
- REPORT: report summarizing and providing visualization of the TST results.
- BUNDLES : Segmented probability bundles.
- TRACKS: Segmented fascicles as trk files.
- CSV: csv files of the scalar measures.
LESION LOAD MEASURES
- CSV: csv file containing the lesion load measures per fascicle.
LESION LOAD MEASURES REPORT
- REPORT: report summarizing and providing visualization of the lesion load results.
It provides the results of the TST report as well.
Create free account now!
References
dMRI PREPROCESSING
-
DWI gradient table check [Ben Jeurissen et al. 2014
-
Subject motion & Eddy correction: [Andersson and Sotiropoulos 2016]
- Distortion correction: [Jezzard and Balaban 1995].
ACPC alignment:
-
ANTs registration: [Avants et al. 2008]
- MNI152 Atlas: [Grabner et al. 2006]
-
FLIRT registration: [Jenkinson et al. 2002]
dMRI tensor and scalar measures
-
MRtrix tools: [Tournier et al. 2012]
-
Tensor estimation: [Veraart et al. 2013]
-
Scalar measures: [Basser et al. 1994], [Westin et al. 1997] ("Geometrical diffusion measures for MRI from tensor basis analysis". Proc Intl Soc Mag Reson Med).
T1 segmentation and cortical labels
-
ANTs script: [Tustison et al. 2014]
-
ANTs registration: [Avants et al. 2008]
-
Atropos tissue segmentation: [Avants et al. 2011]
-
DiRECT cortical thickness: [Das et al. 2009]
-
OASIS-30 tissue priors: [Landman and Warfield 2012] ("MICCAI 2012 workshop on multi-atlas labeling." Medical image computing and computer-assisted intervention conference)
-
DKT40 Mindboggle 101 Atlas: [Klein and Tourville, 2012]
-
AAL Atlas: [Tzourio-Mazoyer et al. 2002], [Rolls et al. 2015]
-
MNI152 Atlas: [Grabner et al. 2006]
MASK_ALIGNMENT
- ANTs registration: [Avants et al. 2008]
Diffusion MODEL
-
msmt_5tt [Ben Jeurisen et al. 2014]
-
fod2dec: Two posters [T. Dhollander et al., 2015] [T. Dhollander et al. 2015]
TRACTOGRAPHY
-
iFOD1 or SD_STREAM: [J-Donald Tournier et al. 2012]
-
iFOD2 poster [J-Donald Tournier et al. 2010]
-
Tensor_Det [Basser, P.J. et al. 2000]
-
Tensor_Prob [Jones, D. et al. 2008]
-
act, -backtrack, -seed_gmwmi [Robert E. Smith et al. 2012]
