Lesion Filling
Applies a lesion-filling algorithm to a T1 image, following the lesions defined by a binary mask. The algorithm reduces the contrast of the lesions voxels, replacing them with values derived from the surrounding healthy tissue. It is intended to improve the quality of further analysis that might be applied to the data, such as morphology studies. This tool also applies resampling to isometric resolution and AC-PC alignment.
Three possible lession-filling algorithms are available:
- Proprietary algorithm (Mint): including modal estimation of values around the lesion, adding simulated noise and blurring effects.
- ANTs: 2D-based, works slice by slice More Info .
Lesion Filling output example
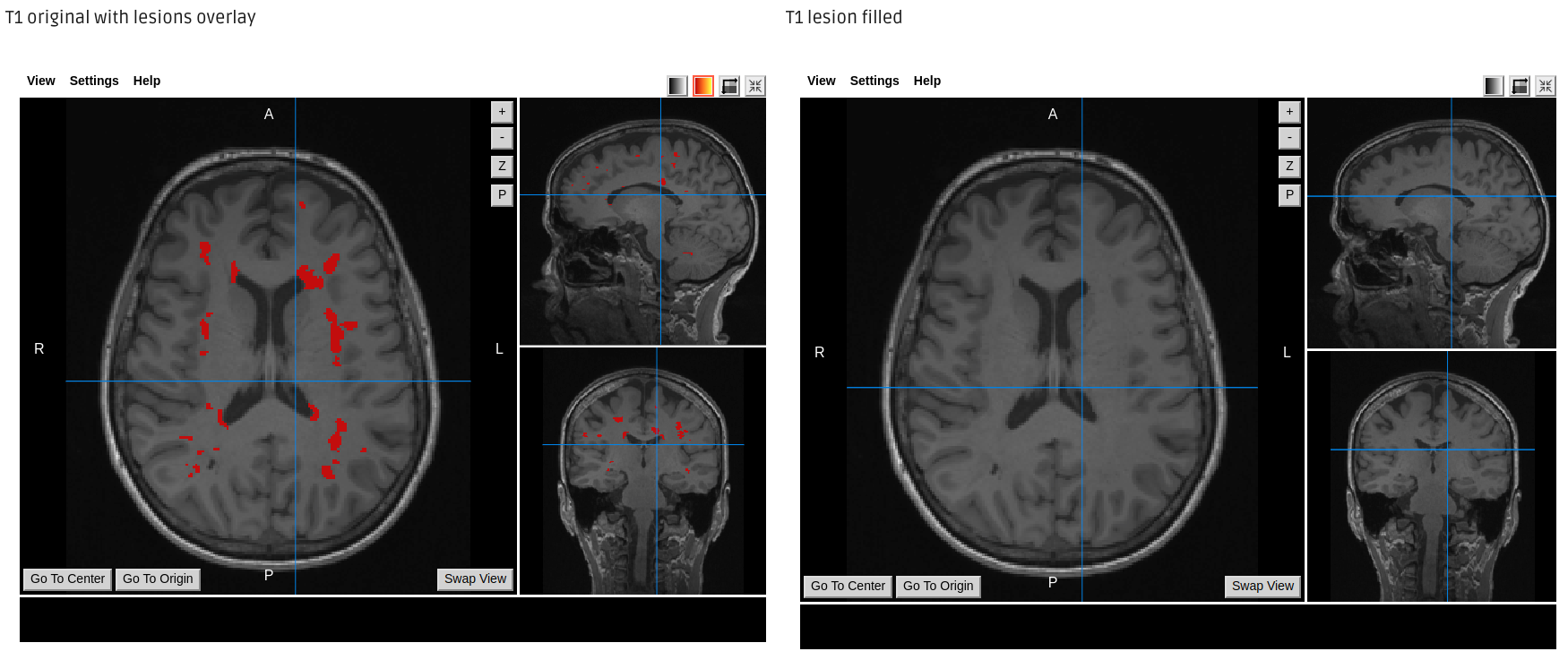
Required inputs:
- T1: anatomical 3D image to be lesion-filled.
Isotropic resolution recommended
Must be labeled as 'T1' modality - Lesion mask: binary image identifying lesion voxels
Must be same resolution as T1 and co-registered
Must be labeled as 'lesions'.
Minimum input requirements:
- T1 Image must contain full skull (not be brain extracted).
- For optimal result reliability, isotropic resolution is recommended
- Recommended resolution: 1 mm isotropic.
- Minimum reliable resolution: 2mm isotropic.
- Lesion mask image must be the same resolution as T1 and co-registered.
Settings:
- Resampling
Choose the desired output isotropic resolution in mm. A 0 value will resample to the highest resolution dimension of the input data
- Isotropic resolution (mm) (float)
- Default: 1
- Isotropic resolution (mm) (float)
- Alignment
- AC-PC alignment (checkbox)
- Default: unchecked
- AC-PC alignment (checkbox)
- Lesion filling
- Lesion filling tool (drop-down selection)
Tool to perform the lesion filling.- QMENTA (Default)
- ANTs
- FSL
- Lesion filling tool (drop-down selection)
Outputs:
- Images:
- T1_acpc_filled.nii.gz: lesion-filled output T1 image.
- T1_original.nii.gz: input T1 image
- lesions.nii.gz: input lesion mask
Typical execution time:
- 5 minutes.
References:
- ANTs registration: Avants et al. 2008
- Lesion-filling:
- ANTs lesion filling: unpublished .
- FSL lesion-filling: Battaglini et al. 2011
Create free account now!
